Structure, allosterics, interactions and function of Hsp70
Chaperones
Erik R.P. Zuiderweg, Ph.D.
Professor of
Biological Chemistry
 Hsp70
Chaperones
Hsp70
Chaperones
Hsp70 proteins mediate trafficking, folding and refolding of proteins in
all known cellular structures. Damage to these proteins is fatal;
overexpression of these proteins is observed in stressed cells, such as in
cancerous tissues. As such Hsp70's are a target for the treatment of
especially breast cancer. These Heat Shock Proteins are 70 kDa, and have a
three domain structure: nucleotide-binding domain (45 kDa), substrate binding
domain (15 kDa) and C-terminal domain (10 kDa). The chaperone helps the
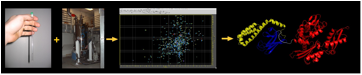
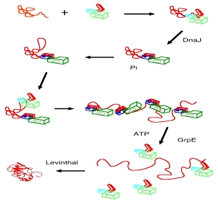
refolding of proteins by binding and release cycles (see pic) driven by
an allosteric mechanism. We use high field NMR in solution (600 -
900 MHz) to study its structure, dynamics and interaction as a function of
substrate, nucleotides and co-chaperones. We are
working on full-length chaperone constructs in order to decipher the allosteric
mechanism (see pic) and to help develop drugs (together with the Jason
Gestwicki lab) to suppress its activity as an aid in Alzheimer therapy.
Summary of our
work in Hsp70 chaperones
Dynamics
methods
The understanding of protein function is incomplete without
considering entropy, that is, dynamics. In enzymes, the active site is
often dynamic to be able to adept to substrate, transition state and product;
for protein complexes, the intermolecular interface sites are
particularly dynamic to accomodate induced fits. Binding processes often
involve perturbation of fast dynamical components, contributing
(sometimes to a dominant extent) to the ligand binding entropy and hence
ligand binding free energy (= affinity). NMR plays an important role in
experimentally measuring dynamics in proteins at time scales ranging from
seconds to pico seconds. Our mission is to develop and apply methods to
describe what the motions actually are.
Summary of our
work in dynamics
Publications of the EZ-group
Varian
sequences in simple programming style