by
Larry "Harris" Taylor, Ph.D.
This material is copyrighted and all
rights retained by the author. This article is made available as a service to
the diving community by the author and may be distributed for any non-commercial
or Not-For-
Profit use.
Go To: Home About "Harris" Articles Slides War Stories Editorials Links Fini
As a biochemist, I am interested in the three-dimensional structure of biologically active molecules and how ligands (drugs) interact with their receptors (site of initiation of biological response). About a decade ago, I was just learning molecular modeling software. (I use a package called SYBYL on an SGI workstation that renders images in true three dimensions). My approach to learning any new-to-me software package is to play with it. So, one rainy Saturday afternoon, as I was beginning to build a library of drug structural coordinate files, I captured the images below. (This was before the days of good image annotation and screen capture, so the technique used was photographing the screen image, which gives text a bit less than crisp.)
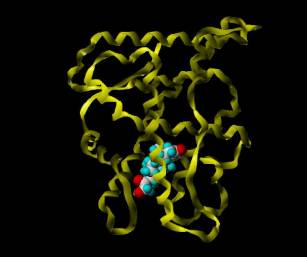
The first image (a steroid within the
MR receptor (involved in the stress
response) is typical of the stuff I do today: This is part of a project I did
for an internet course from Birkbeck College for an Advanced Certificate in
Protein Structure (Modeling
Receptors).

The rest is now decade-old play,
learning various aspects of the software package with a diving motif.










DIVERS ARE SPECIAL
In the realm of biochemistry, each of the amino acids is assigned a letter of the alphabet. This makes it easier for scientists to communicate information about long strings of amino acids (peptides and proteins). One of the things a molecular modeler does is to color amino acids by function and search for "patterns" that might suggest or explain biological activity. The peptide below is hypothetical. It is a theoretical construct, labeled with the proper single letter amino acid code. It is "chemically correct" (for "chemistry geeks" the structure was energy minimized with the SYBYL rendition of the Kollman All Atom Force Field with consideration of Kollman charges.) in that it represents a reasonable geometry for the chosen amino acid sequence.
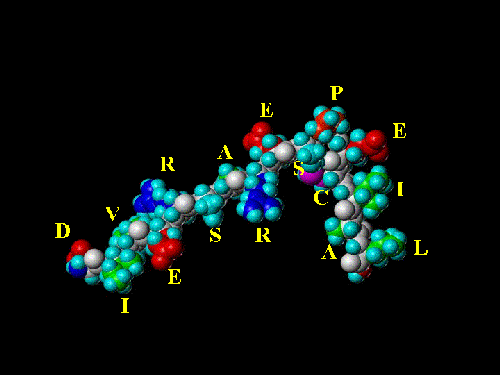
While the molecule is a software construct, I most definitely believe the sentiment expressed in its sequence!
Go To: Home About "Harris" Articles Slides War Stories Editorials Links Fini
About The
Author:
Larry
"Harris" Taylor, Ph.D. is a biochemist and Diving Safety Coordinator at the
University of Michigan. He has authored more than 200 scuba related articles.
His personal dive library (See Alert Diver, Mar/Apr, 1997, p. 54) is considered
one of the best recreational sources of information In North America.
All rights reserved.
Use of these articles for personal or organizational profit is specifically denied.
These articles may be used for not-for-profit diving education