A. Regulation of kinesin motors.
In the absence of cargo, kinesin activity must be tightly controlled to prevent futile ATP hydrolysis and congestion of microtubule tracks. Recent work has demonstrated that autoinhibition may be a general mechanism for the regulation of kinesin motors. The molecular mechanism of autoinhibition has been best studied for Kinesin-1, but members of the kinesin-2 (KIF17/CeOsm-3), kinesin-3 (KIF1A and KIF13/GAKIN), and kinesin-7 (CENP-E) families are also regulated by autoinhibition. As kinesin families evolved by adopting unique non-motor regions for distinct cellular functions, there must have been pressure to ensure the coevolution of inhibitory mechanisms for regulation of the common motor domain.
Molecular mechanisms of Kinesin-1 autoinhibition.
To determine the molecular mechanisms of Kinesin-1 autoinhibition in a cellular context, we utilized fluorescence resonance energy transfer (FRET). Kinesin-1 subunits tagged with donor (mCFP = monomeric cyan FP) and acceptor (mCit = monomeric Citrine, a yellow FP) fluorescent proteins were expressed in COS cells. Three images were obtained from live cells (mCFP, mCit, and FRET) and used to calculate the average FRET efficiency (Eave) of inactive and active motors. Inactive motors localize in a diffuse cytoplasmic pattern whereas active motors can be trapped in a microtubule- bound state using the nonhydrolyzable ATP analogue AMPPNP.
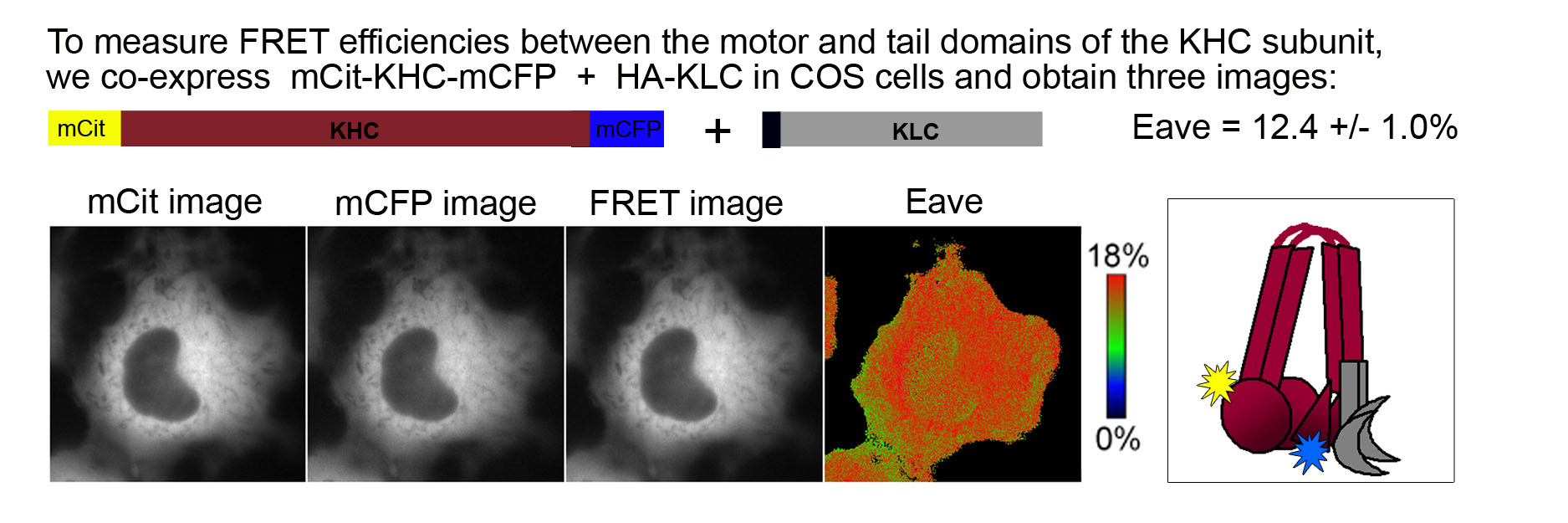
FRET of the inactive Kinesin-1 motor. Fusion of FPs to the N- and C- termini of the KHC subunit indicates the proximity of the motor and tail domains of Kinesin-1 in the inactive state. The fairly high FRET efficiency (Eave = 12.4 ± 1.0%) indicates that the motor and tail domains are in close proximity and thus, that Kinesin-1 is in a folded, autoinhibited state when expressed in mammalian cells.
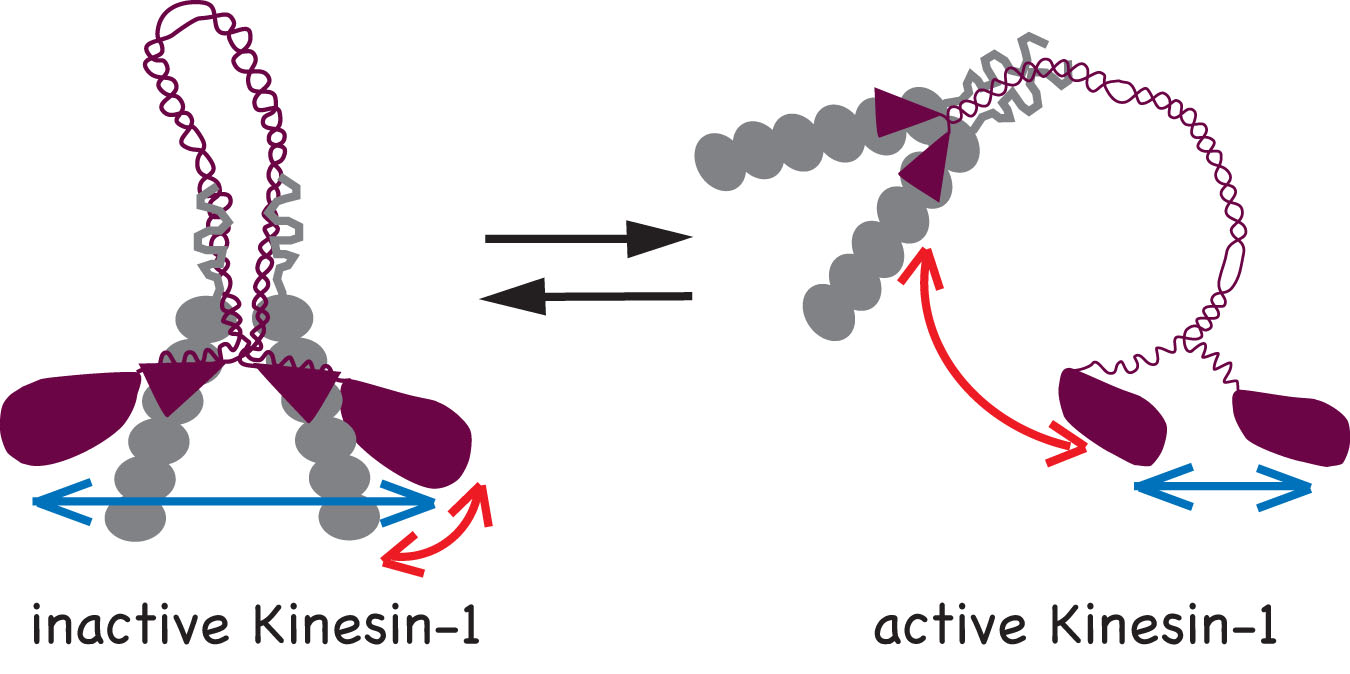
We determined that two mechanisms work to keep Kinesin-1 in an inactive state in the absence of cargo: i) the KHC tail prevents binding of the motor domain to the microtubules and ii) the KLC subunit separates the motor domains so they cannot engage in processive motility. Accordingly, two conformational changes occur upon activation of Kinesin-1: i) a global conformational change where the KHC tail is moved away from the motor domains (red arrows) and ii) a local conformational change that brings the two motor domains closer together (blue arrows).
How is Kinesin-1 activated for microtubule-based transport? The simplest model of Kinesin-1 activation posits that cargo binding to non-motor regions relieves auto-inhibition. However, cargo binding is not sufficient for motor activation. Rather two binding partners, one of KHC (fasciculation and elongation protein zeta (FEZ1)) and one of KLC (JNK-interacting protein 1 (JIP1)), are required to activate Kinesin-1.
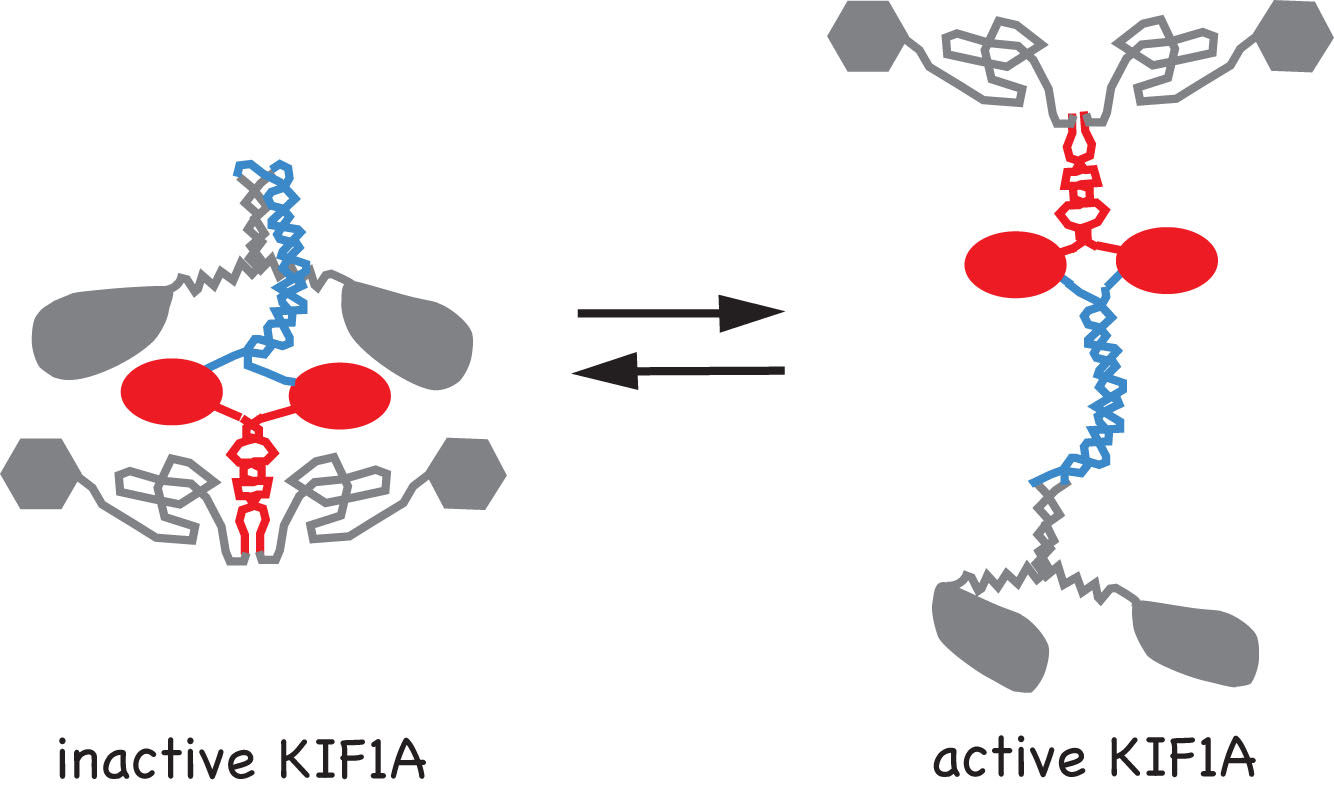
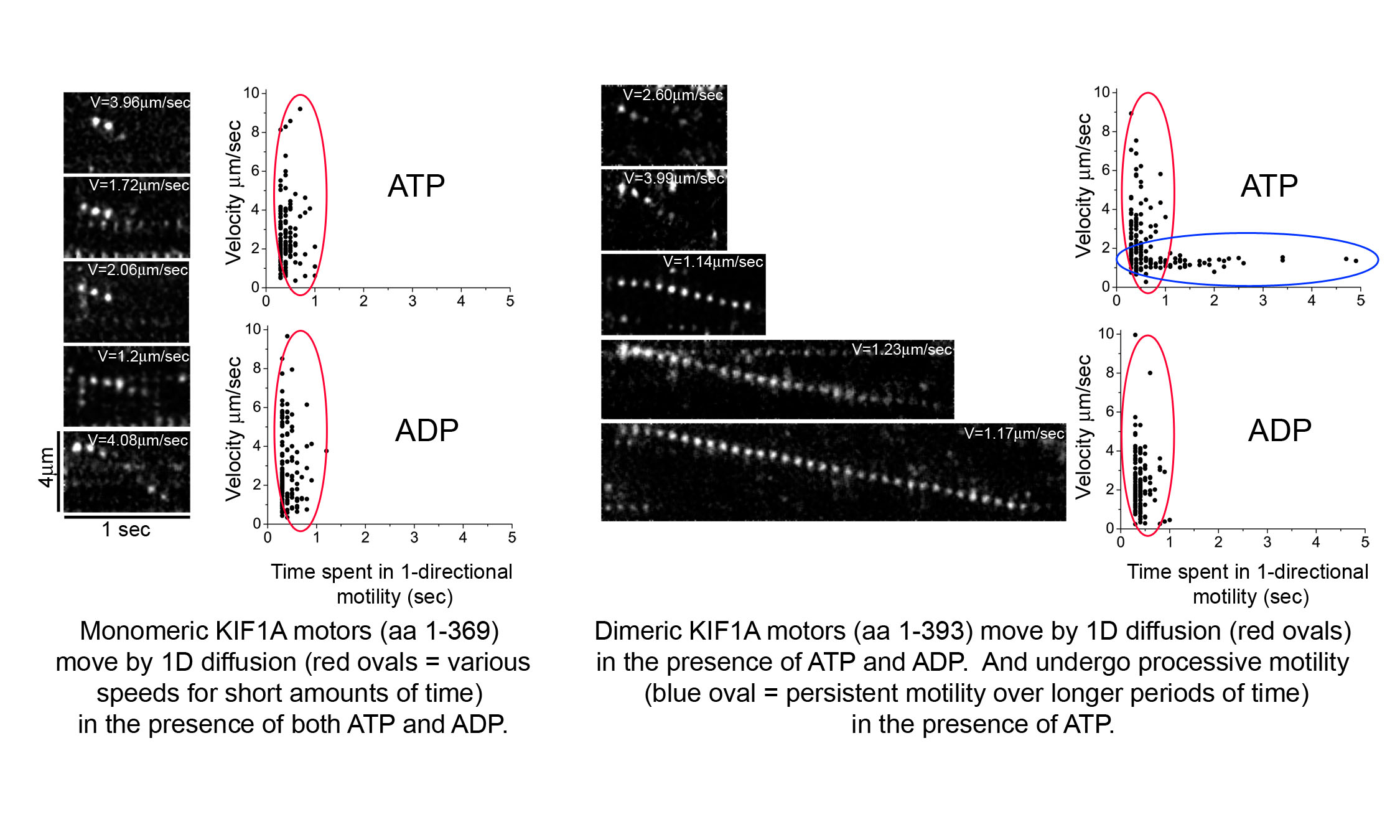
Movies 1-3: Live cell microtubule binding assay of inactive and cargo-activated Kinesin-1. COS cells expressing the indicated Kinesin-1, JIP1 and FEZ1 proteins were transiently permeabilized with Streptolysin O (SLO), washed, and then AMPPNP was added. When expressed alone, KHC-mCit is active and cannot be released from microtubules after addition of AMPPNP (Movie 1). When the Kinesin-1 holoenzyme is recreated by coexpression of KHC-mCit + KLC-mECFP, Kinesin-1 is inactive and cannot be locked on the microtubules (Movie 2). Expression of either JIP1 or FEZ1 alone is not sufficient to activate Kinesin-1 (KHC-mCit + KLC-mECFP) (not shown). However, coexpression of Kinesin-1 with both JIP1 and FEZ1 results in an active Kinesin-1 (KHC-mCit + KLC-mECFP) molecule that can be locked on microtubules upon addition of AMPPNP (Movie 3). Molecular mechanisms of Kinesin-3 autoinhibition.We have also studied the molecular mechanisms that regulate the activity of the kinesin-3 motor KIF1A. We first determined that KIF1A motors expressed in mammalian cells are dimeric motors that are inactive for microtubule binding or motor activity. We then identified two regions of KIF1A that contribute to autoinhibition: i) the FHA domain and subsequent coiled-coil segment (in red) prevent binding of the motor domain to the microtubules and ii) a coiled-coil segment (in blue) prevents processive motility.
We also showed using single molecule imaging of KIF1A motors in in vitro motility assays that dimeric KIF1A motors move along the surface of microtubules by processive motility whereas monomeric motors move by 1D diffusion. Relevant publications:Verhey KJ, Kaul N, and Soppina V. 2011. Kinesin Assembly and Movement in Cells. Ann Rev Biophysics 40: 267-288. PMID 21332353. (REVIEW) Verhey KJ and Hammond JH. 2009. Trraffic control: Regulation of kinesin motors. Nature Reviews Molecular Cell Biology. 10: 765-777. PMID: 19851335. (REVIEW) Hammond JW, Blasius TL, Soppina V, Cai D, and Verhey KJ. 2010. Autoinhibition of the kinesin-2 motor KIF17 via dual intramolecular mechanisms. Journal of Cell Biology 189: 1013-1025. PMID 20530208. Hammond JW, Cai D, Blasius TL, Li Z, Jiang Y, Jih GT, Meyhofer E, and Verhey KJ. 2009. Mammalian kinesin-3 motors are dimeric in vivo and move by processive motility upon release of autoinhibition. PLoS Biology 7: e72. PMID: 19338388. Hammond JW, Griffin K, Jih GT, Stuckey J, and Verhey KJ. 2008. Co-operative versus independent transport of different cargoes by Kinesin-1. Traffic 9: 725-741. PMID: 18266909. Cai D, Hoppe AD, Swanson JA, and Verhey KJ. 2007. Kinesin-1 structural organization and conformational changes revealed by FRET stoichiometry in live cells. Journal of Cell Biology. 176: 51-63. PMID: 17200416. Blasius TL, Cai D, Jih GT, Toret CP, and Verhey KJ. 2007. Two binding partners cooperate to activate the molecular motor kinesin-1. Journal of Cell Biology. 176: 11-17. PMID: 17200414. | |||||||