Research Background


PROFESSIONAL TRAINING AND EXPERIENCE
Education
•Ph.D., Environmental Engineering Science June 1995
California Institute of Technology
Thesis Title: Kinetic, Biochemical, and Genetic Analyses
of the Particulate Methane Monooxygenase
Thesis Advisor: Mary E. Lidstrom
•M.S., Environmental Engineering Science June 1989
California Institute of Technology
•B.S., Civil Engineering with Highest Honors May 1988
The University of Texas at Austin
Positions at the University of Michigan
•Associate Director Aug. 2010 - present
Program in the Environment
College of Literature, Science and Arts
•Associate Professor Jan. 2010 - present
Program in the Environment
College of Literature, Science and Arts
•Graham Fellow for Research Development April 2006 - Dec. 2008
Graham Environmental Sustainability Institute
Office of the Provost
•Study Team Co-Director May 2005 - April 2006
Graham Environmental Sustainability Institute
Office of the Provost
•Associate Professor Jan. 2004 - present
School of Natural Resources and Environment
•Associate Professor June 2001 - present
Department of Civil & Environmental Engineering
College of Engineering
•Assistant Professor Sept. 1995 - June 2001
Department of Civil & Environmental Engineering
College of Engineering
Positions at other institutions or organizations
•Guest Professor March 2009 - June 2009
Department of Environmental Geosciences
The University of Vienna, Austria
•Post-Doctoral Research Fellow Aug. 1994 - July 1995
Department of Biological Sciences
University of Warwick, England
INTRODUCTION TO METHANOTROPHY
APPLICATIONS OF METHANOTROPHY
Pollutant Degradation
Early research has shown that cells expressing sMMO degrade a broader range of substrates than cells expressing pMMO, and in many cases, rapidly degrade these compounds. The range, kinetics, and products of pollutant degradation by cells expressing pMMO, however, had not been carefully examined, nor had the form of MMO predominantly expressed by natural communities been determined.
To address these issues, work in my laboratory has examined the genetics and biochemistry of the pMMO. In this research, we have discovered that methanotrophs expressing pMMO can indeed degrade chlorinated solvents such as trichloroethylene (TCE), a finding previously thought not to be possible. Furthermore, we have found that at least one halogenated hydrocarbon, chloromethane, can serve as a carbon and energy source for methanotrophs when methanotrophs express the pMMO. Recent data published from my laboratory has also shown that in polluted environments, methanotrophs have strong selective pressure to express pMMO in situ as they grow more readily, and can actually degrade more pollutants when expressing pMMO than sMMO (Lee, et al., Applied & Environmental Microbiology, 2006).
Mitigation of Greenhouse Gas Emissions from Landfills
The ability to manipulate methanotrophic activity in situ has important implications for mitigating greenhouse gas emissions from natural and man-made ecosystems, e.g., wetlands and landfills. Methane has a global warming potential approximately 25 times that of CO2. As a result, an emerging topic of research is how to harness the significant potential of aerobic methanotrophic bacteria to control CH4 emissions. As part of a project supported by the Department of Energy, we examined how best to stimulate and monitor methanotrophic activity in landfill cover soils, one of the largest anthropogenic sources of methane. In these environments, although methanotrophic activity can be easily stimulated through the provision of nitrogenous fertilizers, N2O emission rates can and also typically increase. As N2O is approximately 14 times more effective in absorbing infrared radiation than CH4 on a per unit basis, a major outcome of this research was the development of strategies that allowed for the enhancement of methane oxidation while preventing N2O emissions, i.e., we determined how to effectively uncouple CH4 consumption from N2O production such that the emission of both gases can be minimized (Im, et al., Applied Microbiology & Biotechnology, 2010; Lee, et al., Applied Microbiology & Biotechnology, 2009).
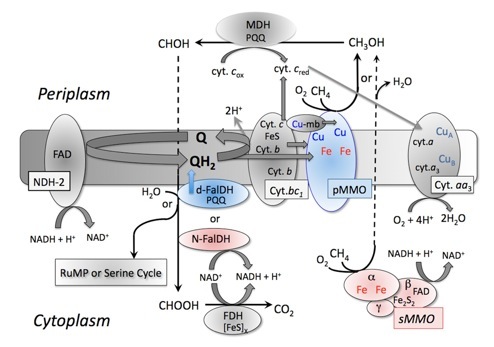
In my current research, I focus on methane-oxidizing bacteria, or methanotrophs. These microorganisms have long been known to be ubiquitous in the environment, found in freshwater sediments, marine sediments, bogs, forest soils, agricultural soils, and aquifers, amongst other locations. Methanotrophs play an important role in the global carbon cycle by consuming methane and have been extensively examined for the biodegradation of priority pollutants. Through co-metabolic reactions catalyzed by methane monooxygenase (MMO), these cells can degrade a wide range of priority pollutants, most notably chlorinated solvents. Two forms of MMO have been found, however, with very different kinetics and substrate ranges. Most methanotrophs constitutively express a membrane-bound, or particulate form of the MMO (pMMO). A small subset of methanotrophs under copper limitation will express a cytoplasmic or soluble MMO (sMMO).
Figure 1. Pathway of methane oxidation in methanotrophs. Proteins showing positive or negative Cu-regulation are shown in blue and red, respectively. Cyt, cytochrome; D-FalDH, dye-linked/quinone-linked formaldehyde dehydrogenase; FDH, formate dehydrogenase; N-FalDH, NAD(P)-linked formaldehyde dehydrogenase; NDH-2, type 2 NADH dehydrogenase; pMMO, membrane-associated or particulate methane monooxygenase; Q, ubiquinone; FAD, flavin adenine dinucleotide; MDH, methanol dehydrogenase; PQQ, pyroloquinoline quinone; sMMO, cytoplasmic or soluble methane monooxygenase; RuMP, ribulose monophosphate. (Semrau, et al., FEMS Microbiology Reviews. 2010).
Despite their critical function in many different ecosystems, as well as controlling methane emissions that may increase due to climate change (methane is a potent greenhouse gas, ~25x more powerful than carbon dioxide in absorbing infrared radiation), the biogeochemical factors that affect their activity and community structure are poorly understood. It is known that copper plays a key role in methanotrophic physiology, but the mechanism used by these microorganisms for copper acquisition was only recently discovered. This compound, analogous to siderophores, termed methanobactin, is the first example of a “copper-siderophore”, or chalkophore. Like siderophores, methanobactin binds many different metals, including mercury, uranium and cobalt, but binds copper preferentially. Furthermore, as found with siderophores, recent data show that different methanotrophs make different forms of methanobactin that have varying metal affinities (Figure 2) (e.g., Krentz, et al., Biochemistry, 2010).
In addition to these studies, we have determined that facultative methanotrophs exist, i.e., methanotrophs that can utilize multi-carbon compounds for growth. Such a finding was unusual as it was initially believed that methanotrophs could only grow on one-carbon compounds such as methane and methanol. Research in my laboratory identified a Methylocystis strain (termed SB2) that could not only grow on methane, but also on acetate and ethanol (Figure 4). When grown on these multi-carbon substrates, we discovered that the pMMO in Methylocystis strain SB2 was constitutively expressed. Such a finding is intriguing as it indicated that strain SB2 could be used for pMMO-mediated pollutant degradation without the need to provide methane, which acts as a competitive inhibitor and also has poor aqueous solubility. We have determined that strain SB2 can indeed degrade a variety of chlorinated hydrocarbons when grown on acetate or ethanol, and this can
FACULTATIVE METHANOTROPHY
CHARACTERIZATION OF METHANOBACTIN
I collaborate with researchers at Iowa State University to investigate the genetics, biochemistry, and biogeochemistry of a novel biogenic metal binding agent, methanobactin, synthesized by methanotrophs. We are currently investigating the ability of methanobactin to bind and solubilize copper in complex systems, e.g., in soils with humic materials that can strongly affect the bioavailability of copper. As part of our initial studies, we have characterized multiple forms of methanobactin, and have discovered that these forms can bind and reduce a wide range of metals, including converting Au(III) to elemental gold (Figure 3). Such discoveries are very intriguing as methanobactin makes well-defined size distributions of gold nanoparticles, which have multiple industrial and medical applications.
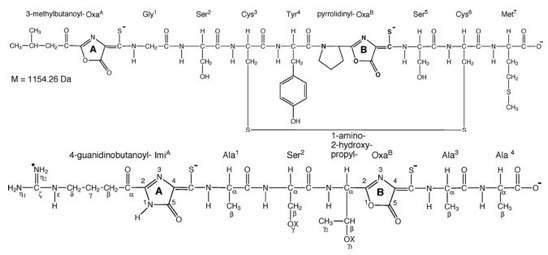
Figure 2. Primary structure of methanobactin from Methylosinus trichosporium OB3b (top) and Methylocystis strain SB2 (bottom).
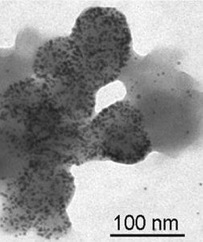

Figure 3. Formation of gold nanoparticles by methanobactin from Methylosinus trichosporium OB3b (Choi, et al., Journal of Inorganic Biochemistry, 2006).
I am pursuing new research to develop better mechanistic and predictive models of the contribution of and interaction between different microbial communities that regulate CH4 emissions from wetland ecosystems in the Arctic region. Specifically, in collaboration with colleagues expert in hydrological and atmospheric modeling,we will: (1) characterize and correlate the distribution, activity, and gene expression of methanotrophs and methanogens to different environmental variables known to affect microbial activity and distribution, e.g., O2, water potential, and temperature; (2) develop a mechanistic, biological/physical model of net CH4 emissions that will couple the dynamics of microbial community structure and activity to the processes of heat-water-gas transfer in soils; (3) integrate the mechanistic model with a larger-scale land-surface model, and; (4) carry out regional assessments of climate change impacts on net CH4 emissions from natural soils and wetlands using Alaska as a “proof-of-concept” domain.
provide alternative bioremediation strategies that may prove promising (Im, et al., Environmental Microbiology Reports, 2011; Im, et al., FEMS Microbiology Letters, 2011; Yoon, et al., Environmental Microbiology Reports, 2011).
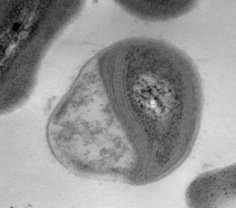
Figure 4. Transmission electron micrograph of the facultative methano-troph, Methylocystis strain SB2 (Im, et al., Environmental Microbiology Reports, 2011).
FUTURE RESEARCH