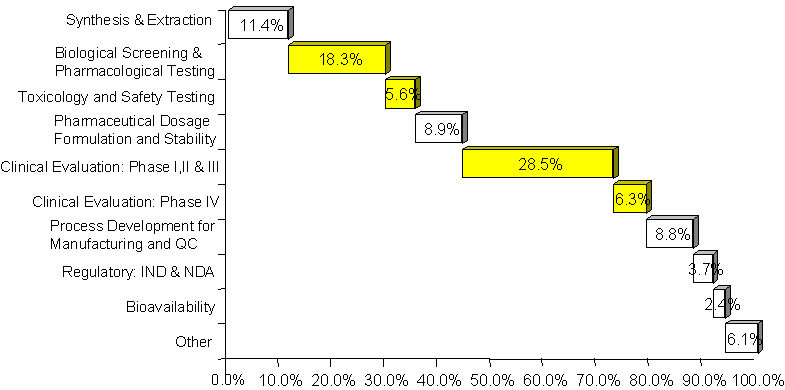Affymetrix Inc.: The GeneChipÒ System
“Ninety-nine percent have NO IDEA how fast
this REVOLUTION IS COMING.”
- Stephen Fodor, Chairman and CEO, Affymetrix Inc.
On New Year’s Eve 1999, Dr. Jean Yeager reflected on her three year career at Affymetrix Inc. Yeager had first joined the Santa Clara, California-based biotechnology firm in 1996 as Vice President of Marketing at a time when the company had virtually no revenue. Through much of 1997 and 1998 she worked closely with visionary CEO Stephen Fodor to begin laying the groundwork for commercializing the firm’s revolutionary technology, the GeneChip® System. During Yeager’s tenure with Affymetrix the company had executed an IPO on the NASDAQ, formed numerous partnerships and alliances, and been the subject of several high-profile patent infringement suits.
Since her promotion to President in September, Yeager had begun to feel the pressure of life at a cutting-edge biotechnology firm. The company had enjoyed no shortage of success to date. Affymetrix had achieved a stock market valuation of nearly $2 billion at a time when Internet stocks seemed to be heavily favored by Wall Street, and the firm was touted by many as the “Microsoft of Medicine.” Affymetrix’s stock price, however, was based on the assumption that the firm would become earnings positive by 2001 and that the GeneChip would become the standard in the rapidly evolving field of genomics. Yeager hoped that Affymetrix would be able to achieve its lofty strategic goals in an industry typified by rapid advances in technology and tremendous sensitivity to ever-changing trends in health care and the legal, social, political, and economic environment.
Affymetrix, Inc.
The seeds for Affymetrix’s innovation in gene chips began under the auspices of Affymax N.V., a Bay Area-biotech firm founded in 1988 by Alejandro Zaffaroni. Zaffaroni was a Uruguayan biochemist known for founding eleven biotechnology companies, ten of which are still in operation. Affymax was developing procedures to automate the process of chemical synthesis, a method that might be used to manufacture millions of potential drug compounds. To date, no organization had developed an ability to perform automated chemical synthesis. Affymax consultant Leigh Read had the idea of imitating manufacturers of semiconductor chips and struck upon the notion of manipulating molecules attached to solid surfaces with beams of light. To accomplish this innovation Zaffaroni and other Affymax scientists believed they needed the expertise of Stephen Fodor, a post-doctoral researcher at UC Berkeley who was
“ …known as a gadgeteer, a big-picture thinker, and a maverick. [Steve] isn’t the sort of guy who’s troubled by [the question] ‘Is it feasible?’…[Affymax] needs Steve’s Fodor’s wonderful hands. Such hands are behind every technology that changes our lives: You find them in the lab night after night, twisting knobs, jotting numbers, soldering wires, hoisting cold cups of coffee. They have a surgeon’s dexterity and an artist’s generative magic. They get ornery contraptions to work. One night about 4 a.m., they flip a switch and something new under the sun starts up like a dream.”[1]
Despite Fodor’s initial unwillingness to leave the academic world for industry, Zaffaroni persuaded him to join the Affymax team, which was working on using chips to manufacture peptides, or short segments of proteins. For various technical reasons, the concept did not produce desired results, so Fodor began to use the prototype chips to manufacture DNA probes (very short strips of DNA) and obtained surprisingly successful results. (See Exhibit 1 and Exhibit 2 for an overview of DNA science.) This innovation worked so well that Affymax spun off Fodor’s team to enable them to focus more on the DNA technology that was ancillary to its core business. In 1993 Affymetrix, Inc. was born with a backing of $50 million in private placements. All future potential of the firm banked on the success of the GeneChip System.
The GeneChipÒ System[2]
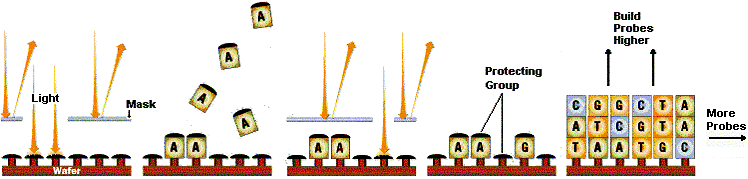
The idea of using silica wafers as the foundation for a system to analyze DNA developed into a product concept called the GeneChipÒ System. This concept is analogous in design to microprocessor chips, except that short DNA probes replace transistors as the base component of the chip. Affymetrix developed a system to manufacture multiple DNA probes simultaneously on a silica wafer. This innovation transformed the traditionally slow serial process of making DNA probes into a streamlined process where millions of reactions could occur in parallel. (See Exhibit 3 and Exhibit 4.) The end product is essentially a matrix of thousands of DNA probes that stand vertically on end on the silica wafer and for which the exact sequence and matrix location of each DNA probe is tracked. (This final product may also be referred to as a probe array, microarray, or DNA chip.)
The microarray requires multiple other components before it can be used as a powerful analytical tool. These additional components include: (1) access to databases containing DNA sequences, (2) stringent manufacturing conditions, (3) chemical solutions and equipment (i.e. a fluidics station) to test biological samples, (3) special laser scanning equipment, and (4) software to analyze the results. These four components together with the GeneChip microarray comprise the GeneChipÒ System. (See Exhibit 5.)
First, information on specific DNA sequences is required to design the DNA probes that will be produced on the chip, and this sequence information must reside on a database that feeds into systems used to manufacture the microarray. DNA sequence information can be pulled from a number of public databases (such as GenBank, the most widely used sequence database) and proprietary sources requiring licensing fees. Next, the microarray is manufactured using Affymetrix’s proprietary photolithographic technique under conditions similar to those required to manufacture microprocessor chips.
Once the chip is manufactured, it is placed in a plastic casing in which the actual chemical reactions will occur. In order to analyze a specific biological sample (such as human blood or cells), special chemical solutions and equipment to automate the procedure are required. When combined under exact conditions, nucleic acids in the biological sample will bind with the DNA probe on the chip only when an exact match exists. (This process is called hybridization.) When an exact match occurs, the match will emit a fluorescent light signal. The complete set of light signals is recorded using a laser scanning device. Finally, software developed by Affymetrix analyzes the optical results recorded from the microarray and compares it with the DNA probe design of the chip to determine the exact gene sequences present in the tested sample.
“The DNA chip is an information-storage device,” says David Singer, Affymetrix’s president and CEO. “It takes a huge amount of information – acquires and processes it – in a way people couldn’t even think of a few years ago.”[3]
The arrival of the GeneChipÒ System as an innovation in DNA analysis was met with wide initial acclaim. Affymetrix’s process for producing the GeneChip is highly reproducible and scalable. This provided potential users of the GeneChip with a standardized process that would be extremely consistent from one use to the next. This consistency was highly regarded throughout the industry as it enabled comparison of results that are acquired not only at different times but also across different geographic sites, two factors inherent in research environments. In addition to consistency, the system is also relatively simple to use, as the entire DNA hybridization reaction is automated. The GeneChip was designed for one-time, disposable use, and this eliminated the hassles of cleaning and decontaminating equipment currently used for DNA analysis. Finally, the system provides a high-density format to analyze huge quantities of DNA sequences. This format allows users the benefits of high-throughput analysis and the associated cost efficiencies of scale economies.
Each microarray, however, is not a universal system and must be redesigned depending on what factors end users want to analyze. For example, microarrays can be designed to detect the presence of genes for a specific disease area (e.g. cystic fibrosis, breast cancer, or Alzheimer’s disease) or a specific species (e.g. mouse or rat genomes that frequently used in research labs), look for small variations in a specific gene (e.g. specific genetic mutations in HIV that cause the virus to become drug resistant), or identify unique similarities and differences in the DNA from different individuals (e.g. paternity testing). The range of analyses that the GeneChip System can perform is limited solely by available gene sequence information. (See Exhibit 6.) Landmark efforts begun in the early 1990s promised to provide the gene sequences needed to analyze wide varieties of diseases and problems.
Corporate Growth
Success followed with Fodor at the helm as scientific director and eventually as president and CEO. Fodor’s team earned not only the 1993 Distinguished Inventor award from the Intellectual Property Owner’s Association, but more importantly won a $31.5 million grant from the U.S. Commerce Department in 1994, the largest grant awarded in that year for the promotion of technology research. By 1994, prototype chips were manufactured in assembly-line fashion, and the Hewlett-Packard Company (H-P) agreed to develop laser scanners to read the GeneChip’s fluorescent output. Within three years H-P was able to develop an extremely fast scanner that provided the capability to read chips containing over 400,000 probes.
After the technical details of the GeneChipÒ system were complete and sourcing was obtained for the fluidics station and scanner, Affymetrix turned its attention to manufacturing the probe arrays. The process for creating the product was very sensitive and required specialized equipment and sophisticated operators. First, Affymetrix hired Ron Verdoorn from Seagate Technology as Executive Vice President of global manufacturing. His background in computer hardware seemed to be an excellent match with the challenges of making silicon-based DNA chips. Second, Affymetrix leased manufacturing space from National Semiconductor, a Bay-Area neighbor that manufacturers etching machines used in semiconductor fabrication. Once technical glitches were resolved, Affymetrix initiated efforts to develop its own full-scale production facilities. (See Exhibit 7.)
The first Affymetrix facility was built in Sonora, CA. As of January 1999, the floorplan was nearly full - six "aligner" machines (the etching equipment for chip fabrication) had been installed. Each aligner had a theoretical capacity of 600 wafers per year;[4] the machines, however, suffered from low yields. Management believed that operator training was one major cause. Another related problem was simply a lack of experience with this equipment. Too much time was spent uncovering problems rather than preventing defects through quality assurance.
Early results were gloomy. Production throughput was equivalent to only 200 wafers per year. Chip yield was equally poor. On average, 50 chips fit on a single wafer (depending on the type of microarrays required). Of these, only 40 emerged as useable.[5] Manufacturing could not meet demand with this yield, and 35% growth in customer demand was expected next year. Feverishly, the firm found a second site and placed orders for more equipment. Due to unexpected delays in the lead time for delivery, the new facility in West Sacramento, CA did not open its doors until mid-1999. By this time Affymetrix had increased the number of aligners at Sonora to ten, maximum capacity for the site. By year’s end each site had improved its yield two-fold, resulting in an annualized throughput of 400 wafers. Chip yield was still a major concern, but microarray design was changing so rapidly that it was difficult to measure yield in a meaningful manner. Due to etching factors current photolithographic masks can only create the GeneChip with distances between DNA probes set at 24 microns. This distance is known as the feature size and dictates the absolute probe density on the chip. Innovations in chip design were further reducing feature size.
The basic layout of the new facility in Sacramento was equivalent to the Sonora facility. The Sacramento plant was built by producing this layout in triplicate, resulting in three identical suites. As of September 1999, however, only one of the suites was equipped with the necessary air handling and other systems required for chip production. Moreover, the company had sufficient trained operators to run two aligners.[6] Management expected to hire two new operators every other month until the end of the year. Despite the production setbacks Affymetrix continued to add customers and was forced to operate both facilities 24 hours a day, 7 days a week, a rate that stressed the manufacturing system and was insufficient to meet demand.
Financial Situation
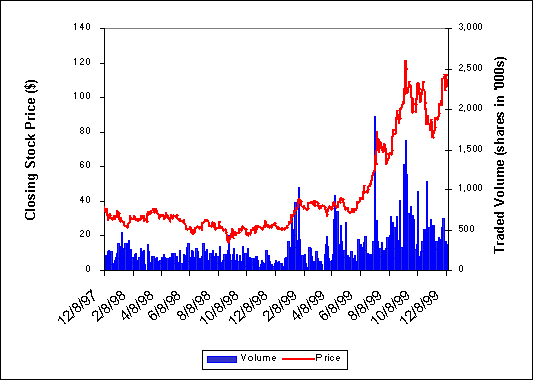
As of January 1995 Affymax owned 65% of Affymetrix when Glaxo Plc, the British pharmaceutical firm, agreed to purchase Affymax for $533 million. The agreement was strategic rather than financially-driven, as Affymax realized it “did not have the capacity to fully exploit [its] drug discovery technology”[7] and thus sought a major partner. In August Affymetrix raised $39 million in private placements, reducing Glaxo’s equity to 47%.[8] On June 6, 1996, Affymetrix launched its initial public offering. The stock was anticipated to open at $15/share, and actual closing price on its first trading day was $17.375 with over 6 million shares issued. The IPO provided the firm with the resources to develop manufacturing and commercialization capabilities as well as additional R&D funding. Since the IPO, the company’s market capitalization has increased dramatically. (See Exhibit 8 - Exhibit 10.) As of September 30, 1999, the conglomerate Glaxo Wellcome owned a 31% equity stake in Affymetrix.[9] (See Exhibit 11 for a history of key corporate milestones.)
The Genomics Revolution
“There’s a revolution going on. You may know it as cloned mice, or the Human Genome Project, or perhaps insect-resistant corn. It’s a revolution with many fronts but one clear quest: unlocking the secrets of genes, the DNA stands that contain the code of life. The implications for humanity are staggering: the prevention of disease, the feeding of the world…The force that’s driving [this revolution] is the rapidly advancing science of genomics [the study of gene structure and function]…If DNA is the software that runs life processes, then scientists want to search for bugs in the programs. Why? As defective software can crash a computer, defective DNA can make you sick. Understand the errors in the code, and you’re on the way to a cure.”[10]
Significant advances in biotechnology and genetics have enabled the decoding of the human genome. The largest effort begun in 1990 is the federally-funded Human Genome Project (HGP), a compilation of scientists striving to determine the exact genetic sequence of all 100,000 human genes within the next several years. This genetic blueprint will be provided through a public database. By the year 2000 experts anticipate that 90% of the human genome will be sequenced, and most of the major genes involved in common diseases will be identified.[11] This project will define the components of human medicine, much like the periodic table does for chemistry, as well as transform approaches to biomedical problems, according to Eric Lander, director of the Whitehead/MIT Center for Genome Research.[12]
These advancements in genetics, molecular biology, and technology have huge implications for companies in the health care and life sciences industries. The ability to predict patient susceptibility to disease, develop more effective drugs in less time, and increase nutritional food content will dramatically change the focus of research and development in these industries. For example, SmithKline Beecham allocated over 25% of its drug discovery programs to genomics work.[13] Furthermore, many companies use mergers, acquisitions, alliances, and licensing agreements to obtain access to critical technology and capabilities from genomics firms.
Alliances and mergers are standard means for pharmaceutical and chemical companies to obtain access to a variety of technology platforms. Recently, a number of scale factors have become increasingly significant in the drug development process, particularly enabling technologies in genomics research. This factor combined with the accelerating costs of R&D as well as the pressure to improve drug pipelines to be competitive and accelerate output has led to new business models and corporate restructuring.[14] (See Exhibit 12.) This phenomenon is not limited to the health care industry; food, nutrition and agriculture companies are also merging with biotech and pharmaceutical firms.[15]
Health care, life sciences and biotech companies have fueled demand for genomics-based products. For practical purposes the genomics market can be segmented into three usage-based groups: research, development, and application.
In research environments, for example, scientists attempt to understand how certain genes manifest disease through mutations. (See Exhibit 13.) Gene expression analysis allows scientists to analyze the correlation of specific DNA sequences and mutations in individuals with and without a specific disease. This process will permit pharmaceutical companies to identify the most promising drug leads, thereby reducing failure rates of drug discovery projects. This development was significant since pharmaceutical firms spent a combined $39 billion in research and development in 1999, and this expense was growing at around 15% per year.[16]
Additionally, a specialized research area on the sources of gene variation between individuals, known as polymorphism analysis, was gaining importance in the industry.
“These variations (in human
gene markers) are what makes each of us unique and more or less susceptible to
specific diseases. Finding and
characterizing a patient’s single nucleotide polymorphisms will usher in the
era of pharmacogenetics and help develop effective drug treatments tailored to
an individual’s particular needs.”[17]
Polymorphisms will allow scientists to determine why individuals treated with the same drug therapy experience different efficacy rates and adverse side effects. “Over 1 million needless hospitalizations and 120,000 mortalities per year could be avoided through pharmacogenetics,” according to Dale R. Pfost, Chairman, President and CEO of Orchid Biocomputer.[18]
Gene expression and polymorphism analysis will also impact the drug development process through targeting better drug leads and improved clinical trials using more effective products. Advanced genomic technologies will also shorten the drug development timeline by allowing thousands of DNA samples to be tested at once instead of in small batches. The combination of enhanced drug pipelines and shorter development cycles will save limited funds and generate higher profits for pharmaceutical companies.
Genomics is also rapidly moving from basic research to practical applications in the medical and life sciences fields, which can be broadly grouped into diagnostics, prognostics, and therapeutics. In the diagnostics arena genetic tests are being created that can improve the matching process of donors for bone marrow transplantation, confirm paternal identity of a child, or diagnose specific strains of viruses, such as HIV and influenza, that mutate frequently. The technology can also identify certain microorganisms in water samples and food borne diseases.
Applications can also be developed to predict whether an individual is prone to certain diseases, thus allowing personalized preventive treatment. Optimists believe that in the future every individual will possess a genetic profile indicating genes that place the individual at risk for disease, thus providing the option of receiving advance therapies. Considering that cardiovascular disease, cancer, Alzheimer’s, diabetes and arthritis account for nearly half of the $1 trillion spent annually on health care in the U.S., the implications of identifying and treating costly and deadly diseases early are mind boggling.[19] As the U.S. population ages and health care insurers continue to focus on cost containment, genomic technologies spell out tremendous opportunities for more effective and less expensive treatments that could prolong life.
Finally, new gene-based therapeutics could be developed to prevent diseases from developing at all. With the identification and sequencing of the genetic code, scientists could genetically engineer DNA to prevent mutant genes from causing disease. This same application could be used in agriculture, food and animals to prevent food spoilage or produce plant-based substitutes for products such as plastics.[20] In addition pharmacogenomics will enable therapies that account for a patient’s individual metabolism and toxicity, thus improving treatment effectiveness and reducing potentially deadly side effects.
If successful, the genomic revolution will create paradigm shifts in health care management, drug development, and life science production.
Commercialization Efforts
Since Affymetrix technologies were still in the early stages of development, the company had only recently commenced product commercialization efforts. To date, the GeneChip system had been sold solely for research use, primarily by pharmaceuticals companies, research institutes and universities. Furthermore, these sales had been almost exclusively for applications in expression monitoring. The Affymetrix customer base was quite small. In 1998 two customers accounted for 36% of total revenues.[21]
Sales
The sales process for enabling research technologies such as the GeneChip had traditionally been a lengthy one. Potential customers were often reluctant to adopt new technologies that would require them to redesign existing research facilities and processes. Introductions from other biotech firms had frequently taken as long as two years to gain momentum.[22] Sales often took the form of long-term supply agreements involving volume-based pricing and collaboration between Affymetrix and customer scientific teams.[23]
Affymetrix generated revenue from three primary sources: GeneChip arrays, hardware and software, and subscription fees. Through partnerships and collaborations the company had begun to make significant progress in commercialization. In September 1999 Affymetrix had an installed base of 170 systems, and was placing new systems at a rate of two per week. (See Exhibit 14 for key customers and collaborators.)
Affymetrix generally offered customers solutions that included broad access to chip arrays, hardware and software, and licenses.[24] A GeneChip System (hardware and software) typically sold for around $160,000. Pricing for chip arrays was based on the number of data points per chip (chip density), chip type (human, mouse, yeast, etc.) chip design and performance. Standard chip arrays were priced around $50 per chip while customized arrays could sell for as much as $850. A typical research customer might purchase 4,000 chip arrays per year. Subscription fees also varied upon the level of access to Affymetrix’s and partner databases and ranged from $250,000 to $5 million per year. (See Exhibit 15 for pricing information by account type.)
To support the sales effort, Affymetrix had increased the size of its sales force considerably. In November 1997 the firm had also entered into a non-exclusive sales agency agreement with Amersham Pharmacia Biotech in an effort to penetrate the biotechnology research segment of the market.[25]
Customers and Commercial Applications
Customers purchased an Affymetrix system to perform one of two types of analysis: gene expression or polymorphism. Expression profiling, the process of analyzing patterns of gene expression, allows researchers to identify the role of a gene in a disease pathway. In late 1999 customer sales for gene expression analysis, including systems, arrays and database subscriptions, represented over 90% of Affymetrix’s revenue stream. Polymorphism analysis, the study of DNA sequence variation had become particularly interesting to customers. In recent years a particular form of variation called Single Nucleotide Polymorphisms (SNPs) had gained significant industry attention. Growth of this market, however, depended on identification and compilation of base DNA sequences, which were then used as “markers” against which to measure genetic variation. Completion of efforts such as the Human Genome Project to determine these sequences would help fuel this growth. In general polymorphism analysis was not currently performed at the scale required to make it cost-effective relative to other options.
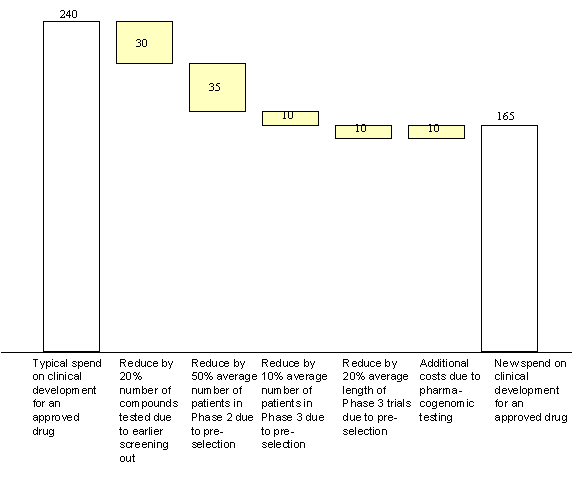
Pharmaceutical customers represented over 70% of Affymetrix’s 1999 revenue base. Customers (such as Merck) employed gene expression analysis in research to identify and validate potential drug targets. This allowed firms to accelerate the costly and time-intensive drug research process where firms typically spent 58% of their R&D budget.[26] (See Exhibit 16.) Sales to pharmaceuticals companies for gene expression analysis had become an important part of Affymetrix’s commercialization program. This revenue stream was expected to grow at an annual rate of approximately 35% for the foreseeable future, even though gene expression had only captured a portion (3%) of the research market to date.[27]
In 1999 large pharmaceutical companies placed enormous value on polymorphism analysis. These customers viewed polymorphism analysis as the next wave of technology, enabling them to isolate key genes associated with disease as well as to tailor medical solutions to individual genetic profiles. Pharmacogenomics would ultimately allow firms to accelerate drug development by allowing them to predict likely adverse reactions more accurately and increase the benefits and efficacy of drugs relative to a specific patient population. This would greatly accelerate the drug development process, particularly in the clinical trials phase. Analysts estimated that in 1999, up to 20% of all development work included some form of pharmacogenomics, and this number was expected to double within 5 years.
The non-pharmaceuticals research market was believed to be 70 – 100% as large as that for pharmaceuticals with equally strong demand. These customers tended to be private research institutes (such as MIT’s Whitehead Institute), government associations (NIH), and academic researchers (University of Michigan). Furthermore, these organizations received funding from multiple sources to acquire Affymetrix’s technology.
“Historically, government organizations had been reluctant to promote Affymetrix’s products broadly due to issues of cost and intellectual property distribution. However, the National Cancer Institute (NCI) announced that it had reached an agreement with Affymetrix under which the NCI would make grant money available to researchers specifically for the purpose of procuring and implementing the Affymetrix technology. In July of 1999 this agreement was expanded to include the National Heart, Lung, and Blood Institute (NHLBI), the National Institute of Allergy and Infectious Diseases (NIAID), the National Institute of Child Health and Human Development (NICHD), and the National Institute of Health Clinical Center (NIHCC).”[28]
These customers currently subscribed to various Affymetrix programs and to date had employed the technology primarily for gene expression work This portion of the business was expected to growth at 35% per year. These customers, however, had also become increasingly interested in performing the type of mapping studies enabled by polymorphism analysis.
Diagnostic customers represented a smaller portion of Affymetrix’s current customer base. Some diagnostics customers were using expression analysis to diagnose patient illnesses. These customers, however, might eventually replace existing processes with expression analysis, including viral and cancer diagnostics and blood testing. Further potential with these customers resided in polymorphism analysis. Industry analysts proposed that diagnostic practices, such as and identity testing could be penetrated with more advanced chip-based solutions. The total clinical and diagnostics market was around $20 billion, though most expected only a portion (35 – 50%) might ultimately be eligible for chip-based solutions. Penetration of this market was expected to progress at a rate similar to that of pharmaceuticals.[29]
Other potential applications for the Gene chip included conventional microbiological research, environmental testing and monitoring and patient dosage optimization (through metabolic profiling). Life sciences companies in particular had become increasingly focused on genomics as a research and development tool for enhanced food products, and such firms spent a combined $26 billion on R&D in 1998.[30]
Through genomics doctors could also help prevent patient adverse reactions to drugs – a major cost to the health care industry. Yet perhaps the greatest implications for chip technology lay in the distant future. Through genome profiling, it was believed that eventual prognostic uses for the GeneChip might immerge. Many in the field felt that, by 2010, testing would establish a genetic profile for all individuals. This profile would suggest the probability of an individual contracting a particular illness (such as cancer). Therefore, preventative treatments could be administered in order to mitigate or avoid the negative consequences of such an ailment.[31]
Competitors
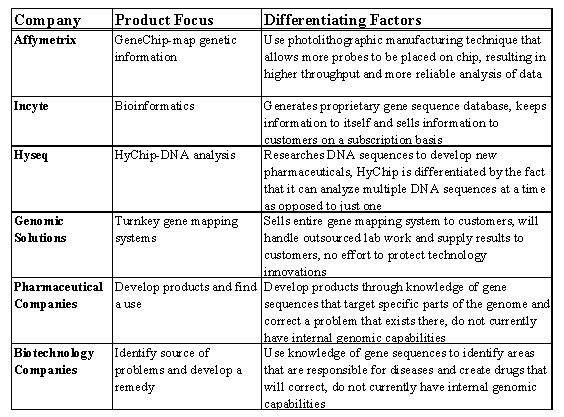
Affymetrix was concerned about its diverse competitor base. Not only did they have to consider direct competitors specializing in genomics but also indirect competitors who were major players in the pharmaceutical and life science fields. (See Exhibit 17.)
Incyte Pharmaceuticals
A former Affymetrix partner, Incyte has taken a slightly different approach in genomics. It has focused on building a library of genetic information on human, animal, plant, and microorganism species. Access to this database is then provided to clients for a subscription fee. Bioinformatics, as it is termed in the industry, allows Incyte to maintain a proprietary database while still taking part in efforts by the pharmaceutical and biotechnology industries to develop new drugs. Other services provided by Incyte include microarray-based gene expression and DNA cloning services and DNA screening.
Incyte had established partnerships with large pharmaceutical companies. It is currently involved in a joint venture with SmithKline Beecham to use genetic data in disease tests. Additionally, Pfizer owns approximately 5% of Incyte and is working with the company to enlarge its database system. These alliances, however have not helped the company’s profitability. While it was recently in the black, declines in subscriptions and negative patent rulings have led Incyte’s earnings to turn negative.[32]
Hyseq Inc.
Hyseq’s core business is to develop biopharmaceuticals using rare genes discovered in-house. Hyseq believes that these rare genes are the key to many diseases and has thus far used them successfully to generate products for cardiovascular, blood-based, and immunological diseases.[33] Hyseq collaborates with partners to develop diagnostic and pharmaceutical potential products. Once the candidates are approved, then the partnership commercializes the products together. The main benefit for a partner is to use Hyseq technology to increase the number of products in its development pipeline. Current Hyseq partners include PE Corporation (owner of PCR technology), Chiron, Kirin, and the University of California-San Francisco.[34]
Hyseq, along with PE Corporation, has developed the HyChip system to perform DNA sequencing and apply results in diagnostic and research settings. Hyseq has provided the research necessary to develop the system, whereas PE Corporation contributes the development, manufacturing, and marketing assets necessary to take a product commercial. The main differentiating factor of the HyChip system is that is can test for a whole array of DNA targets, whereas competing chips require at least one chip for every gene target. Hyseq permits select partners to use the HyChip system before commercial launch. To date, Hyseq has not had a quarter with positive earnings.
Genomic Solutions
Headquartered in Ann Arbor, Michigan, Genomic Solutions has recently opened another office in the UK in an effort to drive global growth. This private company has grown through mergers and acquisitions and is funded by $25 million of venture capital.[35] By utilizing results of a nation-wide effort to map the human genome, Genomic Solutions focused on becoming a one-stop shop for generating new medications.
Genomic Solutions offers turnkey systems to its customers and can supply everything from robotics, biochips, and imaging systems to the software required to run a gene mapping facility. Genomic Solutions also offers laboratory services for cost-constrained customers. Firms can then use services to analyze genes, DNA, and proteins. For companies interested in developing protein-targeting drugs, Genomic Solutions can also provide the requisite applications.
Genomic Solutions has developed an extensive client list that includes most large pharmaceutical companies. The firm has also cultivated relationships with such prominent research universities as Michigan, MIT, Harvard, Berkley, and Johns Hopkins, in addition to the U.S. government and international institutions such as the Max Planck Institute in Germany.[36] Since Genomic Solutions is privately owned, specific profit numbers for the firm are not known, but annual sales are estimated to be in the range of $5-10 million. Genomic Solutions is certainly in a growth stage, and an IPO is expected in the near future.
Other Competitors
When considering competitors for Affymetrix, it is important not to overlook large pharmaceutical and biotechnology firms. While these firms might not work specifically in gene mapping, they have heavily invested in this field and have the ability to be major players. Additionally, pharmaceutical firms that compete heavily in the diagnostics arena can be considered direct competitors of Affymetrix and its gene chip technology.
Furthermore, the biotechnology industry is laden with all forms of non-exclusive alliances. Any firm with which Affymetrix forms a partnership (such as for manufacturing, marketing, or distribution) could become a major competitor within six months. All of the big players in these industries (such as Amgen, Pfizer, Glaxo Wellcome, Monsanto, Eli Lilly, and Bayer) are hedging their bets by developing relationships with as many genomics firms as possible. The result is two-fold. First, large firms are in a position to benefit, regardless of who survives the development phase of genomics companies. Second, these pharmaceutical and biotechnology companies can learn from genomics companies to develop competencies to strike out on their own. Further, tremendous value can be gained by in-sourcing genomics operations. Along these lines, ten pharmaceutical companies have formed a consortium with the mission to internalize all genomics research and development. If successful, results from the consortium will enable pharmaceutical companies to terminate partnerships with fledgling genomics firms. Ultimately, companies that survive the first five years of genomics industry growth develop core capabilities in this area and use them to get products quickly to the marketplace.
Emerging Competitors/Technologies
The GeneChip System is an exciting and revolutionary technology, but Affymetrix is concerned about other technologies that might leapfrog its product during the early stages of commercial launch. Unlikely partners such as Monsanto and IBM were quickly forming alliances to develop new databases and products that would surpass current technology. A major concern for Affymetrix was whether or not it could get to market in time to beat these newly forming competitors or whether its technology would be outdated as soon as it was commercialized. If Affymetrix could launch its products, did it have the critical mass to compete with larger firms? Scientists found themselves facing a problem that continuously plagues all high technology companies: the increasing pace of incremental and disruptive innovations. (See Exhibit 18.)
Pressing Issues
Litigation
As of the third quarter of 1999, Affymetrix was involved in high-profile intellectual property lawsuits with three major competitors: Incyte Pharmaceuticals, HySeq, and Oxford Gene Technologies. These litigations caused expenditures of approximately $10 million annually in legal counsel. Specific case details are as follows:
Incyte Pharmaceuticals: Affymetrix filed a patent infringement suit against Incyte on January 8, 1998. When Incyte acquired Synteni, another biochip competitor in December 1997, Incyte acquired technology that Affymetrix believed was covered under its own patents. As of August 1999, this lawsuit was still pending. Incyte had already suffer from negative patent case rulings and would incur further significant consequences by losing the case, as microarray-related business generated 7-10% of revenues for the firm.[37]
Hyseq: Hyseq filed suit against Affymetrix in late 1997 for infringement of multiple sequencing-by-hybridization (SBH) patents. Affymetrix claims it does not use SBH technology, but complexities of intellectual property law may force the litigation to drag on for at least one more year. Adding to the entanglement, Affymetrix sued Hyseq in August 1998 on issues pertaining to gene chip technology. The outcome of both suits could seriously affect both companies futures.
Oxford Gene Technologies (OGT): OGT was co-founded by Ed Southern, a scientist who pioneered initial efforts in DNA hybridization. (The traditional method of hybridizing DNA is known as the Southern blotting technique.) As such, OGT owns all of the initial patents on DNA hybridization. Initially, Affymetrix tried to license the technology from OGT but eventually got around the license by partnering with Becton-Coulter, a firm to whom OGT had previously extended a license. In June 1999 OGT filed suit against Affymetrix for patent infringement. It is unclear how long it will take to settle this case, but the lawsuit will continue to be a financial burden on Affymetrix’s legal resources for several years.
Manufacturing
Affymetrix faced critical manufacturing constraints. At the end of 1999, new customers were being added weekly, but production could not meet demand. "Build-out" lead times for acquiring equipment and complimentary systems was long and unpredictable. The equipment required highly skilled labor, but people with these skills were difficult to find. At the same time chip designers were continuously crafting chips with greater probe array density, and sales and marketing required manufacturing to customize chips to meet customers' application needs.
The rapid pace of change with this technology made a moving target for the manufacturing organization. Affymetrix's goal was to reach a capacity of 200,000 chips by the end of 2000, but this capacity may still not enable the firm to meet demand. The process is scalable, but Affymetrix still had not mastered medium-scale operations.
Partners/Alliances
Affymetrix had to seriously evaluate its current partnerships. On one hand, the right alliance could provide the firm with marketing and manufacturing resources that it lacked. On the other hand, any partner could take what it had learned from their relationship and become a competitor very quickly. Affymetrix also had to consider that most of its current partners were also working with its closest competitors. It was unclear how the firm should proceed in this regard.
Social Issues
While scientific advances in genomics are altering the health care, life sciences and biotechnology industries, they are also having a profound effect on politics, religion and ethics. The power to detect diseases early, prevent their occurrence, and genetically engineer “super” species raises serious questions about the potential use and misuse of genetic information. For example, moral hazard issues might surface if health insurers and employers had access to information about individuals at-risk for expensive diseases and disabling conditions. Furthermore, the genomics revolution might prove to be a victim of its own success; if scientists can pinpoint and manipulate genetic information, should this be an acceptable practice in society? Although scientific discovery is often perceived as an advancement, worries about discrimination, loss of insurance coverage, loss of employment, and a supernatural human race provide plenty of reasons not to initiate this practice.[38]
Future Challenges
Jean Yeager knew that Affymetrix’s future was uncertain. The GeneChip System had only been used in research applications thus far and might not be viable for commercial use. Significant investments in manufacturing, bioinformatics, software, and quality control systems were necessary for the company to lead the genomics arena on a larger scale. Even with secured funding, Affymetrix still had to undergo the regulatory processes required to launch commercial products. Assuming the firm was able to pass these gateways, there was no guarantee that customers would accept its products. If it were able to launch successful product lines, how long would it be before competitors copied or surpassed its technology? Finally, Affymetrix felt pressure from Wall Street to start generating earnings that would support its lofty stock price.
All of these issues boiled down to three basic points. First, is it in Affymetrix’s best interest to transform itself from a primarily research based company to a commercial company? If so, can Affymetrix continue to secure the funding that is necessary to launch its technologies into the marketplace? Finally, how can the firm protect its products from the patent infringements and disruptive innovations that are a symptom of its industry long enough to secure a sound customer base?
Jean Yeager pondered these thoughts as she watched the ball drop in Times Square, signaling the start of a new millennium. She wondered, would Affymetrix be a revolutionary company in this new genomics era, or would it repeat history and have a fate similar to its parent, Affymax?
Exhibit 1: Overview of DNA and Genomics
All the genetic information needed to form and sustain life is contained in deoxyribonucleic acid molecules, or DNA. Every living creature (including viruses, bacteria, fungi, plants, and animals) use this universal language and DNA instructions are contained in almost every cell.
DNA is composed of four chemicals called nucleotide bases that bind together in a unique way:
Adenine (A) Cytosine (C)
Thymine (T) Guanine (G)
Base A may bind only with base T; base C may bind only with base G.
The bases line up next to each other forming long, slender strands. These strands are similar to one set of teeth for a zipper. Two long strands "zip" together when the bases in first strand bind with the bases in the second strand. These links are called base pairs. As the two strands come together to form a series of base pairs, the "zipper" spirals like a twisting ladder. This formation is called a double helix. The order of base pairs in the double helix is called a DNA sequence. A DNA sequence has typically 150 million base pairs.
Genes are specific groups of base pairs within a DNA strand. Genes vary in length (typically several thousand base pairs) and a DNA strand contains many genes.
Sequences of DNA are contained in structures called chromosomes within the nucleus of a cell. The complete set of chromosomes that map out instructions for an organism is called a genome. The human genome, for example, contains 23 pairs of chromosomes holding 60,000-100,000 genes that collectively have around 3 billion base pairs.
Exhibit 2: Visual Representation of a DNA Molecule
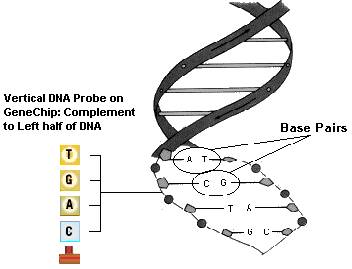
Exhibit 3: Definition of a DNA Microarray
A DNA microarray is an orderly, linear arrangement of nucleotide base "probes". The series of chemical probes in a microarray simulates one half of a DNA "zipper". Probe sequences are designed and constructed to match known (or suspected) genes - nucleotide base sequences within a single DNA strand. When denatured sample DNA is exposed to the probes, sequences of bases from the sample bind to complementary sequences on the microarray.
Typically, multiple microarrays are created for a genetic test, each with a different base sequence (to test for different genes). The location of each array on the test medium is catalogued. After the sample is exposed to the set of probe arrays and hybridization occurs, the arrays are scanned to identify what matches were found in the sample.
The use of microarrays transforms a serial testing procedure (one base sequence at a time) into a massively parallel process where thousands of reactions (tests) can occur simultaneously. Relative to traditional "wet lab" techniques, this is a fast, systematic, reliable process for finding and identifying a large number of genes and gene variations within sample DNA.
Historically, rapid robot spotting was the best technique for creating large numbers of parallel microarrays. This automated the process of affixing a horizontal sequence of nucleotide bases to a solid surface (such as glass) - and improved process precision. In the early 1990s, a new method for synthesizing multiple vertical probe arrays in situ (on chip) was discovered by a team of award-winning inventors at Affymax N.V.. Their discovery led to the Affymetrix GeneChipÒ.
Exhibit 4: The Affymetrix Gene Chip

DNA probe array housed on disposable silica-based gene chips (center) enclosed in plastic casing.
Exhibit 5: The Affymetrix GeneChipÒ System
The Affymetrix GeneChipÒ System consists of tools for acquiring, analyzing, and managing genetic information using DNA microarrays.[39]
· Reagents for targeting and labeling specific nucleic acids.
· Fluidics station to administer the reagents and the test sample onto the probe array.
· Hybridization oven to secure the sample binding to nucleotide bases on the probe array.
· Laser scanner that reads the probe array to identify genetic matches.
· Software that analyzes and manages the genetic information collected from the scanner.

|
Fluidics Station |
Laser Scanner |
Analytical Software |
Affymetrix uses known genetic sequences to design probe arrays that test for specific genetic codes (either a standard set for common genes or custom microarrays for unique research). Using these designs arrays of oligonucleotide probes are synthesized and immobilized onto a silicon wafer. The array is exposed to labeled sample DNA at the fluidics station and hybridized in an "oven." Using the scanner, complementary sequences from the sample are found by locating fluorescent indicators that signal where a match was made. (Probes that have perfectly matched targets generally produce stronger signals than probes with mismatches do.) After the test, the probe array (chip and casing) is then disposed. Included software stores the results of the tests and assists in the analysis of results.
Affymetrix improved its technology at a very rapid rate. The firm learned to make chip features smaller and could therefor fit more microarrays on each wafer. Chip size continued to decrease linearly, while probe density increased exponentially. As of early 1996, Affymetrix designed and produced chips that could measure expressions of 100 different genes. In late 1996, expressions of 1,500 different genes could be performed on a single chip. At the end of 1997, chip capacity was increased to almost 400,000 probes - equivalent to 16,000 expressions.[40]
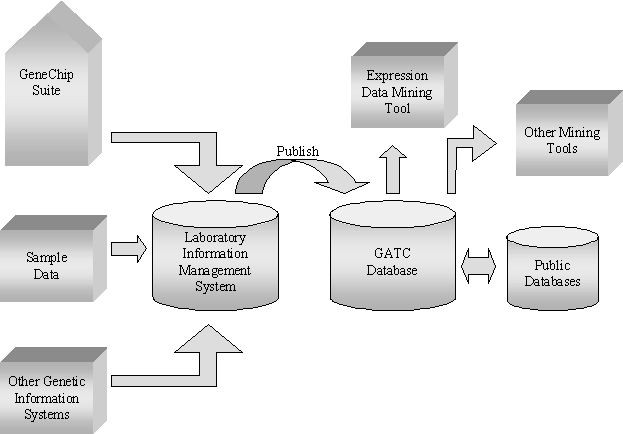
Exhibit 6: The Affymetrix Data System[41]